On This page |
On Related Pages |
|
Install the following dependencies:
EngineVersion" key, newest releases may have issues): See https://downloads.mysql.com/archives/community/ and select the same version as used in production. As an alternative the official docker image https://hub.docker.com/_/mysql can be used to start an instance. Mac user should Launching MySQL with Docker to start an instance through docker compose.
explicit_defaults_for_timestamp is checkedSee Developer Tools for links to the installers.
Make sure you have the Xcode Command Line Tools, which you can install with
xcode-select --install |
I recommend using Homebrew to install Maven and Git. You can also install Eclipse and MySQLWorkbench with Homebrew if desired.
brew install maven git brew tap homebrew/cask brew install eclipse-ide mysqlworkbench |
Using Homebrew to install the JDK or MySQL should be done with caution, because can be difficult to configure Homebrew to use older versions of this software (which Synapse may require as new versions of the JDK and MySQL are released). You can access binaries for the older versions of Java and MySQL with an Oracle account (Google should lead you to these downloads easily).
Your ~/.bash_profile (Note: will be ~/.zshenv if you are on MacOS Catalina or later) should look something like this (where version numbers may differ slightly, and locations may differ entirely if you installed your software differently; check your machine to make sure these folders exist):
export JAVA_HOME=/Library/Java/JavaVirtualMachines/jdk11.jdk/Contents/Home/
export M2_HOME=/usr/local/Cellar/maven/3.5.3/libexec
export M2=$M2_HOME/bin
# The following may not be necessary:
export PATH=${PATH}:/usr/local/mysql/bin # To fix MySQL installation after removing a previous installation via Homebrew |
Using the steps above, you can configure your MySQL server with MySQLWorkbench. You may need to create a file at /etc/my.cnf to edit the configuration and set explicit_defaults_for_timestamp.
Though it might be dated, consider checking out Macintosh Bootstrap Tips for additional reference.
Fork the Sage-Bionetworks Synapse-Repository-Services repository into your own GitHub account: https://help.github.com/articles/fork-a-repo
Make sure you do not have any spaces in the path of your development directory, the parent directory for PLFM and SWC. For example: C:\cygwin64\home\Njand\Sage Bionetworks\Synapse-Repository-Services needs to become: C:\cygwin64\home\Njand\SageBionetworks\Synapse-Repository-Services
Check out everything
git clone https://github.com/[YOUR GITHUB NAME]/Synapse-Repository-Services.git |
Setup the Sage Bionetwork Synapse-Repository-Services repository to be the "upstream" remote for your clone
# change into local clone directory cd Synapse-Repository-Services # set Sage-Bionetwork's Synapse-Repository-Services as remote called "upstream" git remote add upstream https://github.com/Sage-Bionetworks/Synapse-Repository-Services |
Download your forked repo's develop branch, then pull in changes from the central Sage-Bionetwork's Synapse-Repository-Services develop branch
# bring origin's develop branch down locally git checkout -b develop remotes/origin/develop # fetch and merge changes from the Sage-Bionetworks repo into your local clone git fetch upstream git merge upstream/develop |
Note: this is NOT how you should update your local repo in your regular workflow. For that see the Git Workflow page.
devCreate a MySQL user with your named dev[YourUsername] with a password of platform.
create user 'dev[YourUsername]'@'%' identified BY 'platform'; grant all on *.* to 'dev[YourUsername]'@'%' with grant option; |
Create a MySQL schema named dev[YourUsername] and grant permissions to your user.
create database `dev[YourUsername]`; # This might not be needed anymore grant all on `dev[YourUsername]`.* to 'dev[YourUsername]'@'%'; |
Create a schema for the tables feature named dev[YourUsername]tables
create database `dev[YourUsername]tables`; # This might not be needed anymore grant all on `dev[YourUsername]tables`.* to 'dev[YourUsername]'@'%'; |
This allows legacy MySQL stored functions to work. It may not be necessary in the future if all MySQL functions are updated to MySQL 8 standards.
set global log_bin_trust_function_creators=1; |
Note: Use set PERSIST in order to keep the configuration parameter between MySQL restarts (e.g. if the MySQL service is not setup to start automatically). Alternatively, use the Options Files to set this checkbox (under the Logging tab, under Binlog Options).
This allows queries with ORDER BY/GROUP BY clauses where the SELECT list does not match the ORDER BY/GROUP BY clause.
SET GLOBAL sql_mode=(SELECT REPLACE(@@sql_mode,'ONLY_FULL_GROUP_BY','')); |
stackbuilder-dev-[YourUsername] and copy settings from stackbuilder-repo-develop-build. Then click OK
" -Dorg.sagebionetworks.instance=YourUsername"\ 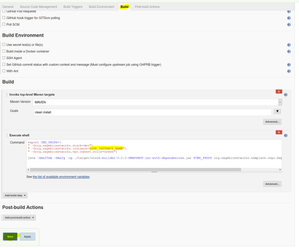
This may be represented as a String Parameter: " -Dorg.sagebionetworks.instance=${INSTANCE}"\, in that case, change the value of the INSTANCE parameter to YourUsername
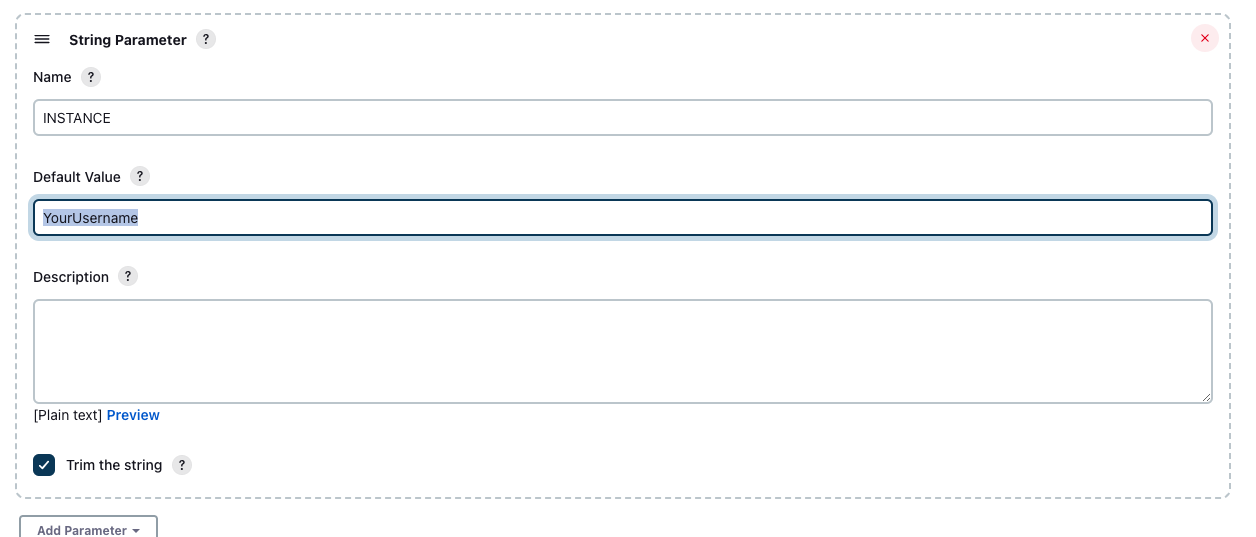
/.m2/settings.xmlUse this settings.xml as your template
<settings xmlns="http://maven.apache.org/SETTINGS/1.0.0"
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
xsi:schemaLocation="http://maven.apache.org/SETTINGS/1.0.0
http://maven.apache.org/xsd/settings-1.0.0.xsd">
<localRepository/>
<interactiveMode/>
<usePluginRegistry/>
<offline/>
<pluginGroups/>
<servers/>
<mirrors/>
<proxies/>
<profiles>
<profile>
<id>dev-environment</id>
<activation>
<activeByDefault>true</activeByDefault>
</activation>
<properties>
<org.sagebionetworks.stackEncryptionKey>20c832f5c262b9d228c721c190567ae0</org.sagebionetworks.stackEncryptionKey>
<org.sagebionetworks.developer>YourUsername</org.sagebionetworks.developer>
<org.sagebionetworks.stack.instance>YourUsername</org.sagebionetworks.stack.instance>
<org.sagebionetworks.stack.iam.id>yourAwsDevIamId</org.sagebionetworks.stack.iam.id>
<org.sagebionetworks.stack.iam.key>yourAwsDevIamKey</org.sagebionetworks.stack.iam.key>
<org.sagebionetworks.google.cloud.enabled>true</org.sagebionetworks.google.cloud.enabled>
<!-- If using not using Google Cloud, set the above setting to false. The following org.sagebionetworks.google.cloud.* settings may be omitted -->
<org.sagebionetworks.google.cloud.client.id>googleCloudDevAccountId</org.sagebionetworks.google.cloud.client.id>
<org.sagebionetworks.google.cloud.client.email>googleCloudDevAccountEmail</org.sagebionetworks.google.cloud.client.email>
<org.sagebionetworks.google.cloud.key>googleCloudDevAccountPrivateKey</org.sagebionetworks.google.cloud.key>
<org.sagebionetworks.google.cloud.key.id>googleCloudDevAccountPrivateKeyId</org.sagebionetworks.google.cloud.key.id>
<org.sagebionetworks.stack>dev</org.sagebionetworks.stack>
</properties>
</profile>
</profiles>
<activeProfiles/>
</settings> |
org.sagebionetworks.google.cloud.enabled to be false. The other Google Cloud parameters can be omitted.Once your settings file is setup, you can Build everything via `mvn` which will run all the tests and then install any resulting jars and wars in your local maven repository.
cd Synapse-Repository-Services mvn install |
Illegal access: this web application instance has been stopped already. This is normal. Scroll up to see whether the build succeeded or failed. (You may have to scroll through several pages of this error.) mvn clean install after debugging your failed build.mvn clean install -DskipTests from the root project, which will rebuild everything. If necessary, run mvn test from the root to ensure all tests pass.If the integration tests fail, try restarting the computer. This resets the open connections that may have become saturated while building/testing.
target/apidocs/index.html and target/testapidocs/index.html for each packageIntelliJ IDEA is an IDE that can be easier to configure to work with the Synapse build than Eclipse. Follow this section if you want to use it instead of Eclipse. Otherwise, you can skip this section.
When you install IntelliJ IDEA, make sure you also install the Maven plugin (should install by default).
There are two settings that you should change in your preferences before installing. You should enable 'Build Project Automatically', and set your Maven home directory to the location of the Maven installation that you used to successfully build the codebase.


After configuring IntelliJ IDEA, you can import the project by pointing to the folder 'Synapse-Repository-Services' and select Maven from 'Import project from external model'. The default settings can be used to finish importing the project.
After this, you need to configure your CLASSPATH. You can easily do this via right-clicking the following folders; mark /lib-auto-generated/target/auto-generated-pojos/ as Sources Root (or Generated Sources Root), and mark /services/repository/src/main/webapp/WEB-INF/ as Sources Root. You may want to confirm that the other folders (listed below in Get your Eclipse Build working) have been marked as Sources Root, but IntelliJ IDEA should have done this automatically. This should complete your configuration, and you can run a few tests to confirm that your repository works.
Note that every time you re-import the above Maven projects, you will need to mark them again.


Add the generated sources to the following projects. (right click and select Build-Path->Use as Source-Folder). If you don't see those folders, refresh the projects until they appear.
/repository-managers/target/generated-sources/jjtree
If you're missing ParseException, you need to confirm the above are added as source folders without any exclusion filters enabled.
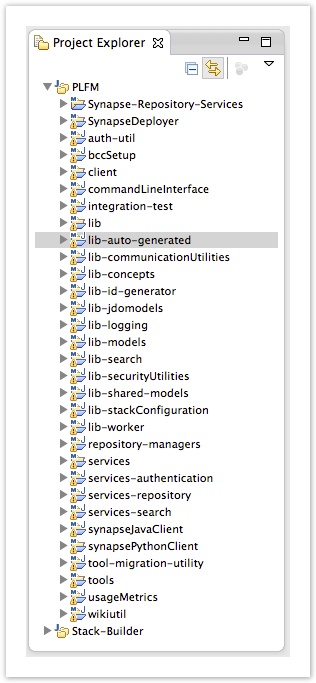
Occasionally, we will make dramatic changes to the PLFM project, like moving or renaming sub-projects. This can cause Eclipse to complain about missing dependencies even after a clean + build (Projects -> Clean... -> "Clean All Projects"). The following steps should help eclipse resolve its dependency problems:
Rebuild all sage artifacts in your local maven repository by running the following from the command line:
mvn clean install -Dmaven.test.skip=true |
Since tests will be skipped this should only take a few minutes. Any build failures in integration-tests can be ignored since the project does not produce any artifacts.
mvn tomcat:run cd integration-tests/ and mvn cargo:run . This should launch a tomcat server on port 8080. When running the application using mvn cargo:run, the context for the API requests will be relative to the version of the repository services. This endpoint will be set to http://localhost:8080/services-repository-${project.version}, where {$project-version} is develop-SNAPSHOT: http://localhost:8080/services-repository-develop-SNAPSHOT/.version and stackInstance of the local project. The web page should load: {"version":"develop-SNAPSHOT","stackInstance":"<your_username>"}To create users for testing, you can use the following script. (Note that the actual username and API key are in fact "migrationAdmin" and "fake". These are only valid on local builds.)
export REPO_ENDPOINT=http://localhost:8080 export ADMIN_USERNAME=migrationAdmin export ADMIN_APIKEY=fake export USERNAME_TO_CREATE=[username] export PASSWORD_TO_CREATE=[password] export EMAIL_TO_CREATE=[email] curl -s https://raw.githubusercontent.com/Sage-Bionetworks/CI-Build-Tools/master/dev-stack/create_user.sh | bash |
To run all the unit tests:
mvn test |
To run just one unit test:
mvn test -Dtest=DatasetControllerTest |
See how to set up a local MySQL
Note that the default user for a locally installed MySQL is root and the default password is the empty string. You only need to use PARAM1 and PARAM2 below if you have configured your local MySQL differently.
To run all the unit tests:
mvn test -DJDBC_CONNECTION_STRING=jdbc:mysql://localhost/test2 [-DPARAM1=theMysqlUser -DPARAM2=theMysqlPassword] |
To run just one unit test:
mvn test -DJDBC_CONNECTION_STRING=jdbc:mysql://localhost/test2 [-DPARAM1=theMysqlUser -DPARAM2=theMysqlPassword] -Dtest=DatasetControllerTest |
Note that these all pass and should continue to do so.
You can use the integration tests, described below, to run a local instance of the service.
Be sure to restore the integration-test/pom.xml file when you are done using remote debugging.
See REST API Documentation Generation
You'll need the following VM Args for the tests or you can put these in your local properties file
-Dorg.sagebionetworks.auth.service.base.url=http://localhost:8080/services-authentication-0.10-SNAPSHOT/auth/v1 -Dorg.sagebionetworks.repository.service.base.url=http://localhost:8080/services-repository-0.10-SNAPSHOT/repo/v1 |
You can do this on Windows using the MySQL Command Line Client. Enter your password and run:
DROP DATABASE <database_name>; CREATE DATABASE <database_name>; |
Prepare and execute a script, local_build.sh, located in the root directory of the git clone like this:
#!/bin/bash
export user=
export m2_cache_parent_folder=
export src_folder=
export org_sagebionetworks_stack_iam_id=
export org_sagebionetworks_stack_iam_key=
export org_sagebionetworks_stackEncryptionKey=
export rds_password=platform
BASEDIR=$(dirname "$0")
${BASEDIR}/docker_build.sh |
This allows running a build without first installing Java, Maven or MySQL. You do need to install Docker first. To execute, run
./local_build.sh |
With a private Jenkins build, you can run unit and integration tests in a controlled environment. In addition, you can integrate your build with your GitHub pull requests to demonstrate that your build has no failing tests (see Push Private Jenkins Build Status to Github as Commit Status).
To access the Jenkins build system you now need /wiki/spaces/IT/pages/352976898. |
To create a private build on Jenkins, create two new jobs
"-Dorg.sagebionetworks.instance=p<user>"export user=p<user>
export org_sagebionetworks_stack_iam_id=
export org_sagebionetworks_stack_iam_key=
export org_sagebionetworks_stackEncryptionKey=
export rds_password=platform
export org_sagebionetworks_repository_database_connection_url=
export org_sagebionetworks_table_cluster_endpoint_0=
/var/lib/jenkins/workspace/${JOB_NAME}/jenkins_build.sh
|
Note, that we add the prefix 'p' to the variable 'user', by convention, to allow local and Jenkins builds to run concurrently without colliding. This is an important aspect since if a local build and a remote build are run concurrently sharing the same stack (e.g. the one that was setup in step 8 of the Synapse Platform Codebase) they might consume shared resources (e.g. queues) leading to race conditions and tests failing. The values for org_sagebionetworks_stack_iam_id, org_sagebionetworks_stack_iam_key, (your IAM ID and Key for your developer account) and org_sagebionetworks_stackEncryptionKey are the same for both local and Jenkins builds. You may point the Jenkins build to the feature branch of your private fork so that it builds whenever you push updates to Github. Also, make sure you change the notification email address to your own in both builds.
org_sagebionetworks_repository_database_connection_url and org_sagebionetworks_table_cluster_endpoint_0 are stack AWS RDS resources created by the stack builder. You can find the endpoints in the RDS console, note that also in this case the database names will contain a reference to the instance created by the stack builder, they should look like something on the lines of:
dev-p<user>-db.blablabla.us-east-1.rds.amazonaws.com
dev-p<user>-table-0.blablabla.us-east-1.rds.amazonaws.com
Now whenever you push changes to the specified fork of your private repository Jenkins will build the stack and run the complete test suite, notifying you of any failures. Once you get a clean build you are ready to create a pull request to the Sage-Bionetworks GitHub repository.
At the end of the local setup + the remote build job you should end up with at least two different stacks in AWS for your account, in the form of dev-<user> and dev-p<user>.
Note: when the stack changes (e.g. new queues are added, new AWS services are setup etc), you might need to run the stack builder again before running your build to avoid having test failing that depends on new or updated resources in AWS. In order to prevent this issue you can setup your private build to always run the stack builder before the build itself:


Now every time the build runs the stack builder will be executed first, make sure that your stack builder job is up to date (e.g. if you use a fork of the Stack Builder, make sure to keep it in sync with the latest upstream develop).
Optionally you might want to parameterize both your stack builder and your build to take in input the branch to run off (e.g. for example if you need a particular stack builder setup for a branch of the repository), you can add parameters to be passed to the stack builder job in the trigger, the parameter value can be prefix with the dollar sign ($) to reference a parameter defined in the current build:

An example of such configuration can be found here: http://build-system-synapse.sagebase.org:8081/job/devpmarco/configure
Note how the job has 3 parameters:
The stack builder job itself is parameterized with a BRANCH parameter (http://build-system-synapse.sagebase.org:8081/job/stackbuilder-dev-marco/configure), the value of this parameter is passed from the upstream job as a "Predefined parameter" (See pic above).
Go to: https://github.com/settings/tokens
Click "Generate new token" button.
Only check "repo:status" and "public_repo".

Create the token and use it in your private Jenkins build's configuration:
export user=p<user>
export org_sagebionetworks_stack_iam_id=
export org_sagebionetworks_stack_iam_key=
export org_sagebionetworks_stackEncryptionKey=
export rds_password=platform
export github_token=<token-created-from-github>
/var/lib/jenkins/workspace/${JOB_NAME}/jenkins_build.sh
|
NOTE: You will need to be a member of the Sage GitHub Organization (https://github.com/orgs/Sage-Bionetworks/teams/synapse-developers) in order to for your token to work. Reach out to IT if you are not yet a member.